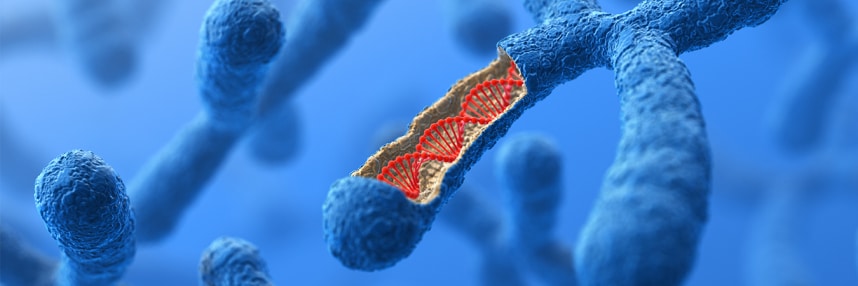
Filling the gaps: sequencing a chromosome
A human chromosome has been sequenced in its entirety for the first time, but why is this breakthrough important for the future of our reference genome?
After two decades of improvements to the human reference genome, a team has published a complete end-to-end sequence of an entire human chromosome for the first time. The study, which used ultra-long-read sequencing to reconstruct the X chromosome, is a significant step forward towards one day creating a complete and accurate human reference genome.
Sequencing the human genome
The first draft of the human genome was published in 2003 as the result of the Human Genome Project (HGP) – an initiative that led to the sequencing of nearly three billion base pairs of DNA.
Since then, sequencing technology has become more advanced, and it is now possible to sequence more quickly and at lower cost. This has led to more ambitious programmes to find out more about the human genome, such as the 100,000 Genomes Project in the UK.
Although the HGP did sequence a significant portion of the genome, there are still gaps in the sequence, and although progress has been made in the intervening period, areas of the genome remain unmapped.
The human reference genome
One of the easiest and most reliable ways to identify differences is to compare to a standard. Scientists have long understood the need for a human reference genome that captures all non-pathogenic human variety; thus making it easier to identify differences which could be pathogenic in new human genome sequences.
Having a reference genome is essential to the practice of bioinformatics. It is used like a template when assembling whole genome or whole exome sequences from sequencer output data, and to find variants that may be linked to health problems or disease.
The accuracy of the reference genome is important because any genomic research or result is only as good as the reference genome used. If the reference genome is inaccurate, then any conclusions drawn by referring to it can also be inaccurate.
Where are we now?
The HGP provided the foundation of today’s reference genome, but it is constantly being improved and kept up to date by the Genome Reference Consortium (GRC), an international body comprising teams at five locations, including the European Bioinformatics Institute and the Wellcome Trust Sanger Institute, both in Cambridge, UK.
The consortium releases revisions to the reference genome regularly, incorporating findings from new research. At the time of publishing, the current version – GRCh38.p13 (Genome Reference Consortium, human, build 38, patch 13) – is the most accurate and complete reference genome for a vertebrate in existence. However, despite many updates, a surprising proportion of the human genome is still missing from the reference.
Where are the gaps?
There are two main areas where gaps exist in the reference genome: the ‘heterochromatic genome’ – sections of the genome where DNA is densely packed around proteins, as well as regions with highly repetitive sequences. In practice, there is a lot of overlap between the two. Neither area was included in the HGP, and so they were missing in the original reference.
Since next-generation sequencing (NGS) became the dominant technology, little progress has been made in these areas. Because NGS works by ‘amplifying’ (making many copies of) small groups of base pairs, these dense and repetitive regions of the genome are extremely difficult to map. It is nearly impossible to piece together these short snippets of sequence data without incorporating too many or too few repeats into the final sequence.
Researchers have only been able to start trying to fill in these gaps since the advent of long-read and ultra-long-read sequencing technologies. Long-read technologies such as Nanopore can read sequences upwards of 10,000 base pairs, with ultra-long-reads of over 100,000 base pairs having been reported. By comparison, Sanger sequencing (the type used in the HGP) and next-generation sequencing can only provide reads of base pairs in the hundreds.
Sequencing a chromosome
In 2018, a team was able to produce a complete sequence of the centromere of a human chromosome using long-read sequencing. The centromere is a region present on every human chromosome essential for keeping dividing chromosomes paired up during cell division and ensuring that each daughter cell receives the correct genetic material.
Centromeres make up around 3% of the genome and are primarily composed of nearly identical repeated sequences. GRCh38 was the first version of the reference genome to include any centromere sequences.
The same team has now followed up with a study that presents the first base-by-base, end-to-end sequence of a human chromosome, including the centromere and other highly repetitive regions. The work is part of the Telomere-to-Telomere (T2T) consortium, which aims to fill in all the gaps in the reference genome.
“We’re starting to find that some of these regions where there were gaps in the reference sequence are actually among the richest for variation in human populations, so we’ve been missing a lot of information that could be important to understanding human biology and disease,” said UC Santa Cruz Genomics Institute assistant research scientist Dr Karen Miga, who led part of the work.
After sequencing the X chromosome, which had 29 gaps in the GRCh38 reference genome, the team now plans to continue its work by sequencing other chromosomes. They will face more challenges, as chromosomes 1 and 9 are known to have repetitive sequences much longer than any found on the X chromosome.
–