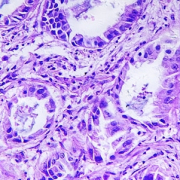
Epigenomic studies spotlight new level of cancer evolution
While genomics sets the scene for cancer to grow, it isn’t the only actor – as shown by two studies that cast a light on the role of epigenomics
A multi-omic level of cancer evolution has been characterised by researchers from the Institute of Cancer Research. In a pair of Nature publications, they highlighted the importance of epigenomic changes to a cancer’s resilience, and how genome-only testing methods may be missing important cancer markers.
Cancer occurs when cells grow and divide uncontrollably. Every cancer is different, yet even within one mass, its cells are not identical: the cells can differ by which genes they express and their susceptibility to treatment.
Both studies examined the evolution of colorectal cancers and tried to understand why cells within a cancer differ in terms of what drives cancer growth and cancer cell survival.
Epigenomic changes are common
In the first of the two studies, researchers looked at 30 patients with bowel cancer. They examined each cancer’s genome and its epigenome. Using this multi-omic approach, they found that changes to the epigenome were common around a cancer’s driver genes. Cancer driver genes are genes that are normally involved in a cell’s division and growth but DNA mutations alter their normal functioning.
Additionally, the epigenomic changes were found to be passed on to the next generation of cells; they were present in cancer cells that had survival advantages over other cells.
- Further learning: What is epigenomics?
Tumours persist due to variation across its cells
Cells within a cancer mass can often be different from each other. The second study took a deep dive into the differences found within tumours. The researchers did this by sampling many different sections of individual tumours to understand how and why the cells of a single cancer can be so diverse.
They found that, while there is variation in the expression of genes within tumours, in most cases this doesn’t alter the ability of cells to survive or grow. However, if the cancer’s environment changes (such as during anti-cancer treatment), then this variation becomes important for its future survival as it may mean that some of its cells survive while others do not. Ultimately, it’s the cancer’s surviving cells that pass on their resistance to treatment to their daughter cells, meaning the cancer, in some form, persists.
“For years our understanding of cancer has focused on genetic mutations which permanently change the DNA code,” said Institute of Cancer Research director and lead author Professor Trevor Graham, “But our research has shown that the way the DNA folds up can change which genes are read without altering the DNA code, and this can be very important in determining how cancers behave.”
How is this being used in practice?
Knowledge around cancer epigenomics and how it fuels cancer growth is already being used in cancer detection. For example, the Galleri multi-cancer test, which we reported as being trialled in the NHS, uses epigenomic markers to detect non-symptomatic cancers. Yet the epigenome is not widely being used in precision cancer treatment. If it was, Professor Graham noted that, “we could, potentially, much more accurately predict which treatments will work best for a particular person’s cancer”.
Tools are available that could make personalised cancer treatment a reality. We previously discussed long-read sequencing approaches, like nanopore, which allow real-time ‘reading’ of DNA to understand the genome, and parts of the epigenome too.
On the studies, Professor Graham noted: “I hope our work will change the way we think about cancer and its treatment.”