Liquid biopsy – a new era in personalised medicine?
Examining the promises and challenges of exciting advances taking place in genetic testing, aiming to improve diagnosis and reduce patient risk
Personalised medicine in the field of cancer treatments is now well established, allowing a more stratified approach to chemotherapy based on sub-groups of patients whose tumours share common genetic make-up. It is unlikely these days that many new drugs will become available without an identifiable associated molecular target, allowing a more rational targeting of therapy rather than a ‘one size fits all’ approach. However, despite the clear benefits of genetic testing to inform stratification of treatments, it is not always simple to achieve. Fiona McDonald considers.
What’s the problem?
Currently, tissue samples for diagnostic testing are obtained by biopsy or surgery, both of which are invasive procedures with potential complications. In addition, for some tumours (for example, in the lung), tissue is not always easily accessible. Monitoring the effectiveness of treatment for the early identification of the development of drug resistance and prompt switching to an alternative drug when necessary requires serial sampling of the tumour’s genomic make-up. Repeated biopsies are associated with morbidity for the patient and can also be costly for healthcare providers.
Biopsies contain only small amounts of tissue, and initial routine histopathology testing may use up all the available material before any DNA can be extracted. The process of tissue fixation can also result in a number of chemical reactions, which lead to the introduction of DNA sequencing artefacts resulting in false positive results. These are just the more practical sides of the problem. A more significant challenge comes from tumour genetic heterogeneity (variation) and also the genomic changes associated with clonal evolution as tumours accumulate mutations that allow them to expand and metastasise.
Heterogeneity exists not only between tumours, but also within them. The spatial distribution of mutations across a tumour means that a biopsy is a sampling exercise at best, such that an important therapeutic target in one part of a tumour can easily be missed. For example, in renal cell tumours it has been shown that only a third of mutations are found throughout the tumour, with the remainder found only in localised areas. Moreover, the genomic make-up of a tumour is not static. Tumours will acquire new mutations as primary tumours and their metastases develop, fuelled by the pressure of anti-cancer therapies, and some of these mutations will be associated with drug resistance. A novel approach is therefore required to overcome these various problems to enable a more accurate approach to personalised treatments.
A new alternative
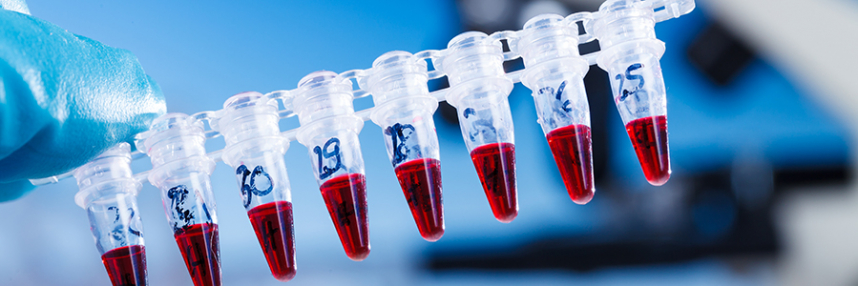
The field of prenatal diagnosis has been revolutionised by non-invasive diagnosis using cell free foetal DNA present in maternal plasma. It has also been recognised for several decades that cell free tumour DNA (cfDNA) can be found in the plasma and serum of patients with a variety of different tumour types. Progress in making use of this information for clinical purposes has been slower than for prenatal diagnosis because of the lack of well-defined targets, but this field has now caught up.
cfDNA originates from cell death and rupture, leading to the release of fragmented DNA into the bloodstream. The DNA fragments released by tumour cells harbour the same genetic changes as the tumour itself, and collectively provide a better representation of the entire tumour landscape (including both primary tumours and their metastases) compared to site specific changes seen in a single biopsy of a tumour site. This has led to the use of the term ’liquid biopsy’.
Analysis of cfDNA is technically challenging, as the tumour DNA fraction is highly variable and has been measured as anything from 0.01% to as much as 90% of the total cell free DNA. However, in a presentation at the American Society of Clinical Oncology (ASCO) in June this year1 it was shown that even when tumour cfDNA makes up less than 0.4% of the total cf DNA, the accuracy of testing were still high. Technological advances have made high accuracy cfDNA analysis possible, notably the methods of droplet digital PCR and next-generation DNA sequencing.
Where are we now?
cfDNA analysis has rapidly moved into the cancer diagnostic arena. One of the initial applications has focused on non-small cell lung cancer (NSCLC), in which 10-15% of tumours contain epidermal growth factor receptor (EGFR) activating mutations which make the tumours susceptible to the EGFR tyrosine kinase inhibitors (TKI) gefitinib and erlotinib. Despite the good initial response to treatment in mutation positive tumours, the majority subsequently acquire TKI resistant mutations. The gold standard for EGFR analysis has been mutation analysis of resected tumours or biopsies followed by serial sampling to monitor the development of resistance. Given the complications from invasive procedures to obtain tissue and the limitations posed by typically small samples, that cfDNA testing for has proved popular.
Initial data, including a meta-analysis of 27 studies2, showed that detection of EGFR mutations in cfDNA had a high diagnostic accuracy, revealing the same genomic make-up seen in the corresponding tumours, and could successfully be used as a prognostic marker. Diagnostic services for EGFR analysis in cfDNA are now available at a number of UK centres and there is already an established external quality assurance scheme. The ASCO presentation2 showed that the results from nearly two thirds of patients with a variety of tumours would make them eligible to join an approved drug trial. It is not surprising that cfDNA analysis is set to make a big impact on personalised medicine in the immediate future.
- Qiu et al. (2014) Cancer Epidemiology and Prevention, 24, 206-212
- www.asco.org/about-asco/press-center/news-releases/liquid-biopsy-may-help-guide-treatment-decisions-advanced
Fiona Macdonald is a consultant clinical scientist based primarily in the West Midlands Regional Genetics Laboratory, and is a fellow of the Royal College of Pathologists. She is an author on more than 80 papers and has also written a variety of book chapters as well as co-authoring two books on the molecular basis of cancer.
–